How Los Alamos Is Learning to Track Disease Outbreaks Around the World
In the 14th century, the Black Death devastated Europe. This bacterial disease killed some 200 million people, almost two-thirds of the Europeans then alive. Preventing the spread of the disease became an urgent public health issue, particularly for ports such as Venice.
So the Venetian Republic took radical action. The government appointed three guardians of public health whose role was to spot ships carrying infected individuals and exclude them from the seaport. This exercise was the first public health measure ever undertaken by a European government. And it later went further by detaining travelers from plague-infected areas for 40 days to prevent the further spread of the disease.
Today, the work of detecting, tracking, and investigating diseases affecting human, animal, and plant health is called biosurveillance. And it is a field that is growing rapidly.
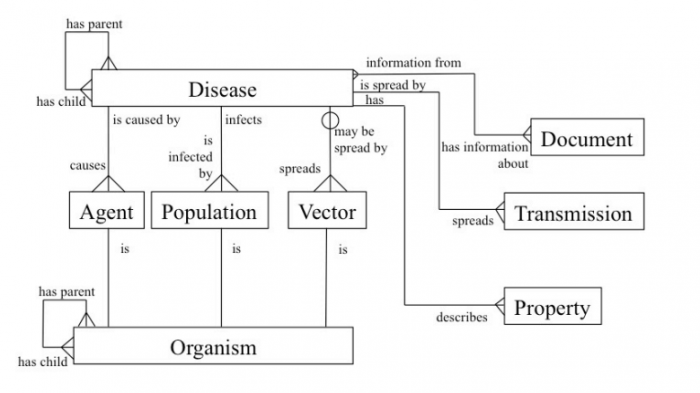
But there is problem. Despite the ancient origins of biosurveillance, there is little agreement over how best to pursue it. Consequently, the field is plagued with difficulties over how to define diseases, their symptoms, infectious agents, their carriers, and so on.
That’s caused no small amount of confusion. Epidemiologists have found to their cost that it is hard to track a disease if nobody agrees on how to describe its characteristics.
Now that looks set to change thanks to the work of Ashlynn Daughton at Los Alamos National Laboratory in New Mexico and few pals who have come up with a new method for describing disease that is designed to bring this disparate field together and gain international traction. Their new system of classification is called the Anthology of Biosurveillance Diseases, and they have set up an online database to support it.
The problems the database is designed to overcome are all too easy to see. One common problem is the confusion that synonyms for diseases can cause. For example, German measles is not a term for measles but for the disease rubella. And rubeola is a synonym not for rubella but for measles. “It was vital to ensure that our ontology capture these synonyms, and others like them, without confusion,” say Daughton and co.
Another part of biosurveillance is to accurately record how a disease spreads. So the new system allows a detailed description of the vectors involved. For example, it can record that dengue is transmitted by the Aedes aegypti mosquito, while chikungunya is spread by the Aedes albopictus mosquito and avian pox by mosquitoes in general.
Some conditions are often described or searched for by their symptoms, so this has to be taken into account as well. And when diseases are notifiable, it must be possible to quickly alert public health bodies.
Daughton and co have developed a database that can be easily interrogated by application program interfaces (APIs) and incorporated into existing biosurveillance tools, of which there are several that lack the features Daughton and co say are so useful.
There are challenges, of course. One is that the entire database has been put together by a team of human experts, a task that has been both tricky and time-consuming. And the task is likely to become more onerous as more diseases are added to the database to cover not just humans but animals and plants as well.
That raises the important question of how the process of keeping the database up-to-date can be improved. “We are interested in insights into ways some of this process can be automated,” say Daughton and co. Clearly there’s a task here for machine learning experts with a few hours to spare.
In the meantime, Daughton and co have made their database available at https://brd.bsvgateway.org/disease/. The government of the Venetian Republic would surely be impressed.
Ref: arxiv.org/abs/1609.05774: A Globally-Applicable Disease Ontology For Biosurveillance; Anthology Of Biosurveillance Diseases (ABD)
Keep Reading
Most Popular
Large language models can do jaw-dropping things. But nobody knows exactly why.
And that's a problem. Figuring it out is one of the biggest scientific puzzles of our time and a crucial step towards controlling more powerful future models.
How scientists traced a mysterious covid case back to six toilets
When wastewater surveillance turns into a hunt for a single infected individual, the ethics get tricky.
The problem with plug-in hybrids? Their drivers.
Plug-in hybrids are often sold as a transition to EVs, but new data from Europe shows we’re still underestimating the emissions they produce.
Stay connected
Get the latest updates from
MIT Technology Review
Discover special offers, top stories, upcoming events, and more.