Terahertz Chip Identifies Short Strands of DNA
One of the more significant practical challenges currently occupying molecular biologists is to find better ways of identifying short strands of DNA. Called oligonucleotides, these strands of nucleotides are hugely useful in processes such as genetic testing, forensics and DNA amplification.
But identifying the strands is a somewhat laboured business. Almost every detection method relies on fluorescent dyes and markers that can be picked up by optical sensors providing a useful but indirect indication of the molecules that are present.
But molecular biologists would like a better system that measures the characteristics of the molecules involved and so provides direct evidence of the sequence of nucleotides. Indeed, various research teams are working on such systems, some with significant success.
Today, Andrey Chernev at St Petersburg Academic University in Russia and a few pals say they have invented an entirely new way of identifying oligonucleotides using terahertz radiation. “Our results demonstrate a new method for label-free, real-time oligonucleotide characterisation,” they say.
An oligonucleotide is a short single-stranded DNA or RNA molecule usually consisting of fewer than a hundred or so bases. The sequence of these bases determines the type of oligonucleotide. So the ideal detection mechanism would reveal this sequence.
Chernev and co’s idea is based on the way these molecules resonate. They say that the sequence of bases in an oligonucleotide determines the way in which the strand resonates at frequencies in the terahertz range. Their idea is to capture a single oligonucleotide in a cavity filled with terahertz waves that stimulates this resonant behaviour.
They begin by producing a signal as close as possible to the resonant mode. By measuring the output from this cavity, they can determine when the input spectra exactly matches the resonant modes of the molecule. That tells them exactly what sort of oligonucleotide they have.
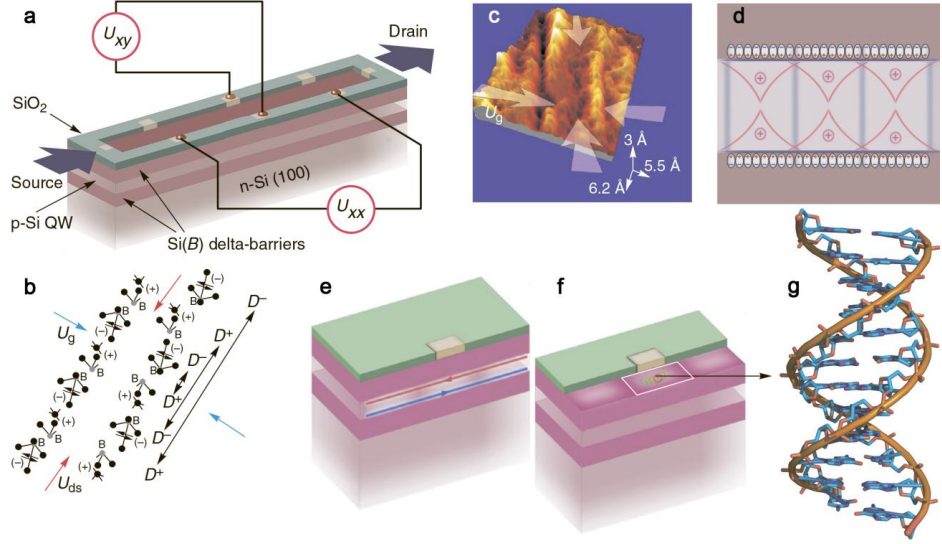
That’s the theory and they have tested it using a device they call a silicon nanosandwich, a quantum well of p-type silicon surrounded by barriers doped with boron. This produces terahertz radiation inside the well where the oligonucleotide is deposited at a concentration that allows a single molecule to enter.
They have tested the device on two oligonucleotides, one that was 50 bases long and the other that was 100 bases long and characterised their unique resonances. That allows them to distinguish between the two oligonucleotides with relative ease at room temperature.
But this is just a first step. What is needed next is an entire library of the unique signatures associated with every oligonucleotide. That should be possible. Chernev and co aim to start by analysing the resonances of each of the monomer and dimer molecules that make up oligonucleotides. The signatures from these should provide a kind of alphabet from which to work out the resonances of more complex polymers.
That is an interesting approach. But this is a crowded area in which lots of good ideas are competing to become the standard technique for identifying the molecules of life quickly and cheaply. Chernev and co will have their work cut out to prove that their method is better than the others that are emerging. But it certainly has potential and so is one to watch for the future.
Ref: http://arxiv.org/abs/1407.6520 : DNA Detection By THz Pumping
Keep Reading
Most Popular
Large language models can do jaw-dropping things. But nobody knows exactly why.
And that's a problem. Figuring it out is one of the biggest scientific puzzles of our time and a crucial step towards controlling more powerful future models.
How scientists traced a mysterious covid case back to six toilets
When wastewater surveillance turns into a hunt for a single infected individual, the ethics get tricky.
The problem with plug-in hybrids? Their drivers.
Plug-in hybrids are often sold as a transition to EVs, but new data from Europe shows we’re still underestimating the emissions they produce.
Stay connected
Get the latest updates from
MIT Technology Review
Discover special offers, top stories, upcoming events, and more.