Device Finds Stray Cancer Cells in Patients’ Blood
Doctors typically diagnose cancer via a biopsy, which can be invasive and expensive. A better way would be to detect telltale tumor cells floating in the bloodstream, but such a test has proved difficult to develop because stray cancer cells are rare, and it’s difficult to separate them from the mélange of cells in circulation.
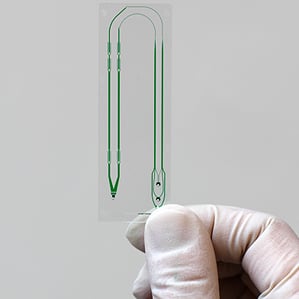
Now researchers from Massachusetts General Hospital and Harvard Medical School say they’ve built a microfluidic device that can quickly grab nearly any type of tumor cell, an advance that may one day lead to simple blood tests for detecting or tracking cancer.
Similar, existing devices—including earlier versions developed by the authors of the current study, which appears in Wednesday’s online issue of Science Translational Medicine—depend on tumor-specific biomarkers on the surface of the cells to pull them out of a blood sample, meaning that a given device won’t work for all cancer types. What’s more, the process of separating the tumor cells from other cell types is generally inefficient and time-consuming. In a given blood sample, circulating tumor cells are rare—there may be only one tumor cell for every billion cells.
The new device is a “substantial step forward from previous microfluidic devices,” says Peter Kuhn, who studies circulating tumor cells at the Scripps Research Institute but was not involved in the study. The device combines existing microfluidic techniques of cell sorting into a single device, he says. The result is that the tumor cells can be pulled out of a blood sample quicker, and without prior knowledge of their molecular characteristics.
Mehmet Toner, director of the BioMicroElectroMechanical Systems Resource Center at MGH, and colleagues report that their latest chip can isolate circulating tumor cells in the blood and that it could help detect all types of cancer. “For our earlier chip, you needed to know something on the surface of the tumor cells,” says Toner. In those devices, a small sample of blood would flow through microfluidic chambers, some of which contained an antibody that grabbed tumor cells. That system also took four to five hours to process a single blood sample. “But for early detection and to make this useful for virtually all cancers, we needed to increase the throughput and to make it [tumor-type] independent,” he says.
Identifying these wandering tumor cells could also help researchers study a cancer’s progression and help doctors track treatments or screen for new cases. By studying the surface proteins or genetic profiles of the cancer cells, doctors and researchers could learn which mutations are present in the cancer and perhaps tailor molecularly targeted treatments accordingly. The authors show that 15 tumor cells were recovered from a blood sample from a prostate cancer patient. The gene expression levels of each cell were studied individually and a mix of mutations was found.
The device developed by Toner’s group combines magnetic labeling of cells and microfluidic sorting to process a sample of blood in about an hour or two. To capture tumor cells regardless of their cancer type, the system first tags white blood cells with magnetic beads that are covered with antibodies that recognize proteins on the surface of the immune cells. The sample is then passed into microfluidic chambers that clear out red blood cells, plasma, and unused free magnetic beads based on their size. Then the device discards the tagged white blood cells using a magnetic field. “In the past, we were focused on tumor cells that we know very little about,” says Toner. “Here, we throw away the cells we know everything about, the blood cells,” he says.
The advantage of the new cell-sorting device over previous attempts is that it successfully brings together multiple technologies already used in the field, such as size separation and magnetic-tag separation, says Gajus Worthington, president and CEO of Fluidigm, a California company that produces microfluidic devices for biomedical research. “The key thing here is the integration, which is crucial to anything related to single-cell work,” he says. All the steps in Toner’s device take place in similar volumes. “If you have to go from one microstep back to macrostep back to microstep, there are losses and complexity, which leads to noise,” says Worthington.
Toner notes that the ultimate goal of identifying circulating tumor cells would be to diagnose patients early. “About 10 percent of cancer patients survive if they are diagnosed late, but almost 90 percent survive if they are diagnosed early,” he says. But whether or not these circulating tumor cells can be found in early-stage patients is not yet clear, says Luis Diaz, an oncologist at Johns Hopkins University School of Medicine, who was not involved in the study. “Early-stage cancers might release very few cells into circulation,” he says. “That’s historically the problem with circulating tumor cells; you can only find them in advanced cancers.”
Keep Reading
Most Popular
Large language models can do jaw-dropping things. But nobody knows exactly why.
And that's a problem. Figuring it out is one of the biggest scientific puzzles of our time and a crucial step towards controlling more powerful future models.
How scientists traced a mysterious covid case back to six toilets
When wastewater surveillance turns into a hunt for a single infected individual, the ethics get tricky.
The problem with plug-in hybrids? Their drivers.
Plug-in hybrids are often sold as a transition to EVs, but new data from Europe shows we’re still underestimating the emissions they produce.
Google DeepMind’s new generative model makes Super Mario–like games from scratch
Genie learns how to control games by watching hours and hours of video. It could help train next-gen robots too.
Stay connected
Get the latest updates from
MIT Technology Review
Discover special offers, top stories, upcoming events, and more.